 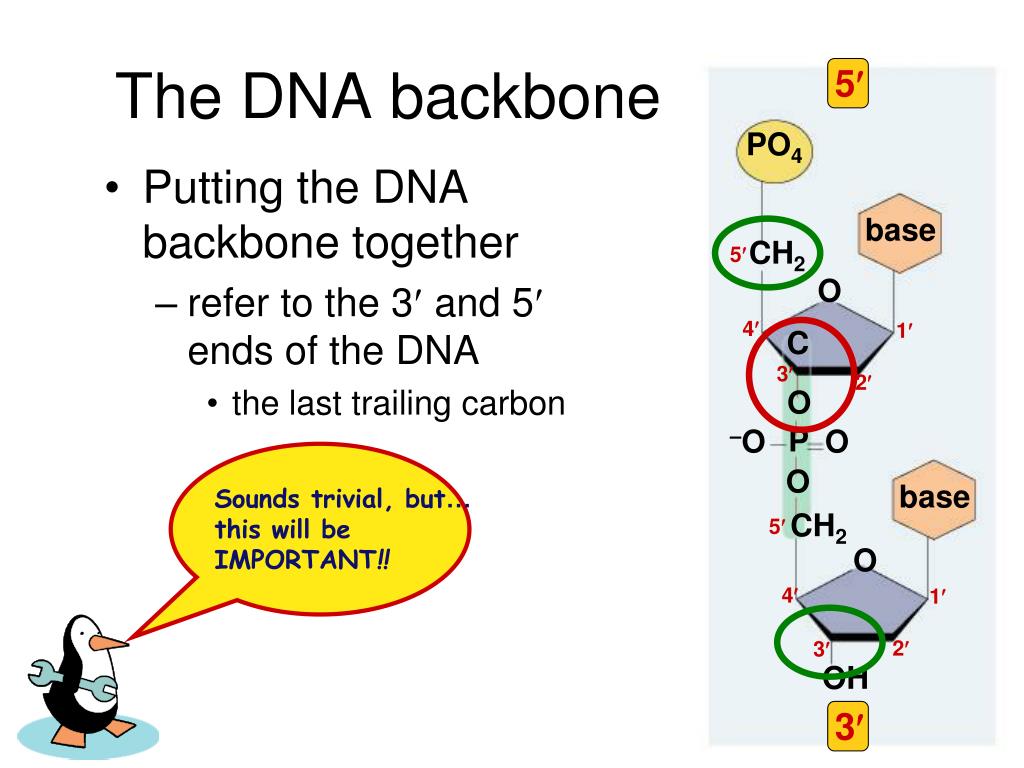
In order for the phosphodiester bond to be formed and the nucleotides to be joined, the tri-phosphate or di-phosphate forms of the nucleotide building blocks are broken apart to give off energy required to drive the enzyme-catalyzed reaction. The negative charge attracts histones, metal cations such as magnesium, and polyamines. Repulsion between these negative charges influences the conformation of the polynucleic acids. Phosphodiesters are negatively charged at pH 7. These saccharide groups are derived from deoxyribose in DNA and ribose in RNA. Specifically, the phosphodiester bond links the 3' carbon atom of one sugar molecule and the 5' carbon atom of another (hence the name, 3', 5' phosphodiester linkage ). 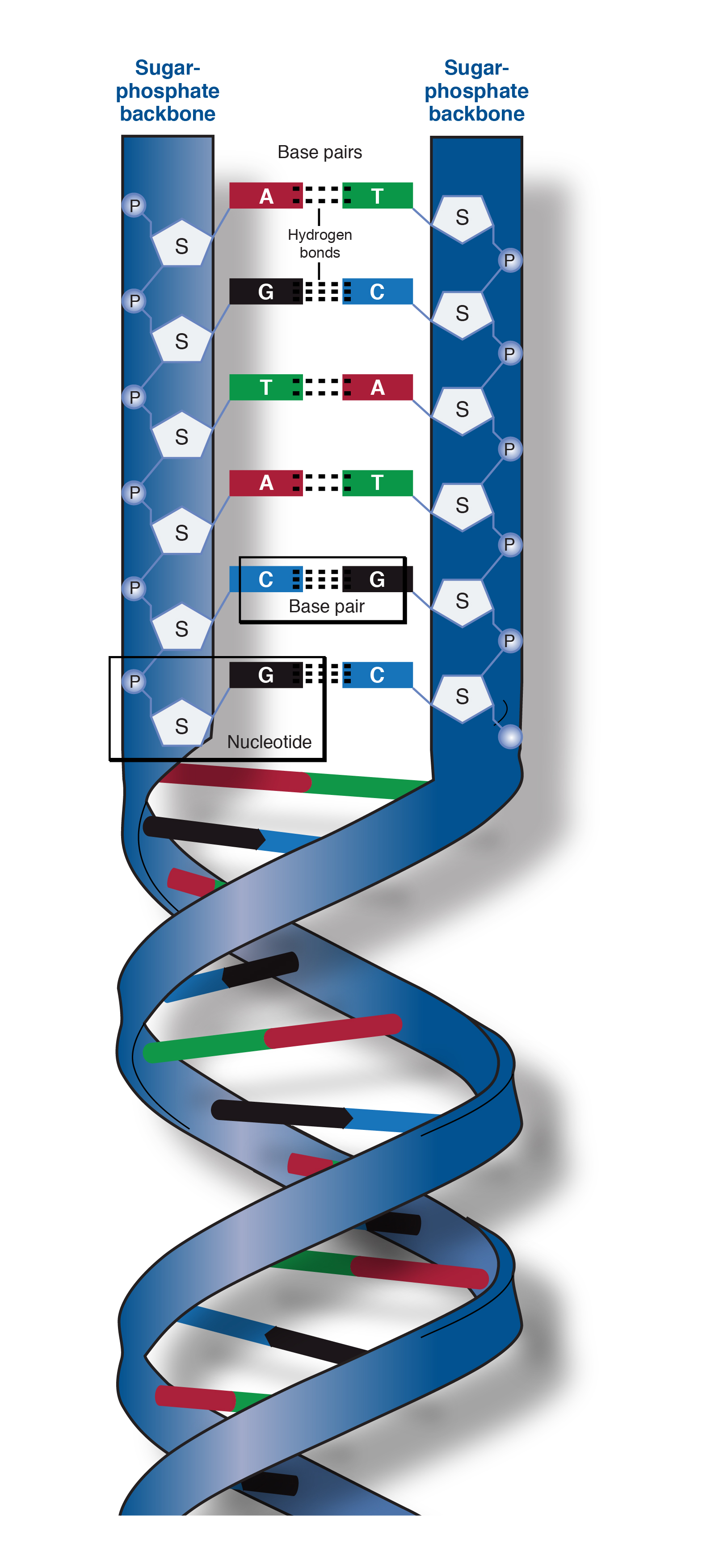
The 3' carbon of one sugar is bonded to the 5' phosphate of the adjacent sugar. The phosphate is attached to the 5' carbon. Phosphodiester bonds make up the backbones of DNA and RNA. Discussion of phosphodiesters is dominated by their prevalence in DNA and RNA, but phosphodiesters occur in other biomolecules, e.g. The "bond" involves this linkage C−O−PO − 2O−C. In chemistry, a phosphodiester bond occurs when exactly two of the hydroxyl groups ( −OH) in phosphoric acid react with hydroxyl groups on other molecules to form two ester bonds. The 5' end has a 5' carbon attached to a phosphate, and the other end, the 3' end, has a 3' carbon attached to a hydroxyl group. Although the graphs represent a compilation of individual base pairs in many duplex structures, the average backbone-base inclination angle based on a single structure is typically of sufficient predictive value for a particular duplex family or class of nucleic acids.Diagram of phosphodiester bonds ( PO 3− 4) between three nucleotides. The tighter distribution of inclination angles with A-form duplexes is apparent in all panels and is not a consequence of the different numbers of observations included in the analysis (29 A-DNA structures and 25 B-DNA structures see the Materials and Methods section), but is likely a manifestation of the more limited conformational flexibility of A-DNA relative to B-DNA. The average inclinations and SDs for A- and B-form duplexes are given above the histograms. The definition of the backbone direction affects the backbone-base pair inclination angle η B: ( A) calculating a BSpline curve through P atoms ( B) vectors connecting P atoms from adjacent residues along the strand ( C) calculating a BSpline curve through C1′ atoms and ( D) vectors connecting C1′ atoms from adjacent residues along the strand. The negative inclination of RNA is consistent with the higher stability of duplexes with 3'- compared to those with 5'-dangling ends.īackbone-base pair inclinations in A-DNA (red) and B-DNA (green) duplexes. Nucleic acid pairing systems with significant backbone-base inclinations can also be expected to display different stabilities depending on which terminus carries unpaired nucleotides. Thus, systems with strongly deviating eta(B) angles will not pair with each other. In addition to providing a rationalization of relative strand polarity with nucleic acids, the backbone-base inclination parameter is also a determinant of cross-pairing. On the other hand, the more or less perpendicular orientation of backbone and bases in DNA allows it to adopt a ps rWC paired duplex. Accordingly, RNA with a backbone-base inclination of -30 degrees, pairs strictly in an aps fashion. This observation illustrates the handicap encountered by nucleic acid systems with inclinations eta(B) that differ significantly from 0 degrees to form a ps rWC paired duplex. 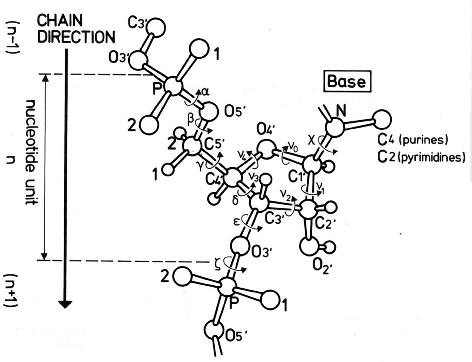
However, all other tested systems are unable to form stable ps duplexes with reverse Watson-Crick (rWC) base pairs. As shown previously, DNA is capable of forming both ps and aps duplexes. The thermodynamic stabilities of the corresponding aps duplexes were also measured. The potential of formation of ps duplexes between complementary 16-mers with eight A and U(T) residues each was investigated with DNA, RNA, 2'-O-methylated RNA, homo-DNA and p-RNA, the ribopyranosyl isomer of RNA. Here, we show its biological significance as a predictive measure for the relative strand polarities (antiparallel, aps, or parallel, ps) in duplexes of DNA, RNA and artificial nucleic acid pairing systems. We have defined a parameter eta(B) that corresponds to the local inclination between sugar-phosphate backbone and base plane in nucleic acid strands. Conversely, the tight coiling of strands conceals that the backbone-base inclinations for A- (DNA and RNA) and B-form (DNA) duplexes differ considerably. This inclination is easily perceived because homo-DNA exhibits only a modest helical twist. The crystal structure of the duplex formed by oligo(2',3'-dideoxy-beta-d-glucopyranosyl)nucleotides (homo-DNA) revealed strongly inclined backbone and base-pair axes. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
